Reliable Extraction of Follow-Up Actions and Timelines from Clinical Notes via Joint Span-Link Modeling and Deterministic Date Normalization
Abstract
Follow-up instructions in outpatient notes often specify actions that must occur at future times, such as imaging, laboratory testing, specialist consultation, or rehabilitation. These instructions are clinically important but are frequently embedded in free text, where the relevant action, the temporal expression, and the association between them may be separated by distractors, section boundaries, or multiple competing instructions. We study the task of extracting structured action-date items from clinical notes, decomposing it into action span detection, time span detection, action-time linking, and date normalization anchored to the visit date.
We present a hybrid neural-symbolic pipeline that combines a shared BioBERT encoder with a BIO tagging head for action and time spans, a span-pair linking head for assigning times to actions, and deterministic normalization of relative dates. On a 2,000-note synthetic outpatient corpus covering five specialties and controlled style variation, the pipeline achieves 0.995 action-span F1, 0.997 time-span F1, 0.980 action-date F1, 0.985 exact date accuracy on matched actions, and 0.53-day mean absolute date error. Under the same held-out split, direct generative baselines show lower action-date F1 and larger date errors. These results support the hypothesis that separating semantic extraction from calendar arithmetic can improve reliability for structured follow-up extraction.
The present study should be read as a controlled proof of concept rather than evidence of deployment readiness. The corpus is synthetic, the results are point estimates from one split, and prospective validation on real de-identified clinical notes remains necessary. We describe the current evidence, the relation to clinical temporal information extraction and LLM-based clinical information extraction, and the experiments required to make the work suitable for a top-tier clinical NLP or biomedical informatics venue.
Keywords: clinical natural language processing; information extraction; temporal expression normalization; relation extraction; BioBERT; follow-up instructions; synthetic clinical data
1. Introduction
Electronic health records contain a large volume of free-text documentation that records clinical plans, diagnostic uncertainty, rationale, and patient-specific instructions. Clinical information extraction has therefore become a central problem in biomedical informatics, with applications spanning cohort discovery, quality measurement, decision support, registry construction, and downstream validation of structured EHR fields [1]. Follow-up instructions are a particularly consequential subset of this information. A note may say that a patient should obtain an MRI in two weeks, repeat laboratory testing in three months, schedule physical therapy within five weeks, or return only if symptoms worsen. Failure to capture such instructions can affect scheduling, care coordination, and auditability.
The technical challenge is not simply named entity recognition. A clinically useful extraction must recover a structured item, such as {"action": "MRI Brain", "period_text": "in 2 weeks", "period_date": "2026-01-24"}, where period_date is computed relative to the encounter date. This requires recognizing action spans, recognizing temporal expressions, linking the correct time to the correct action, and normalizing relative dates. The difficulty increases when notes contain multiple actions, shorthand expressions such as 3 mos or q6mo, historical temporal distractors, and plan sections whose structure varies across specialties and authors.
Prior clinical NLP has established strong foundations for concept extraction and temporal reasoning. Systems such as cTAKES operationalized modular clinical concept extraction for EHR text [2], while the i2b2 temporal challenge, THYME corpus, and Clinical TempEval tasks formalized event and time extraction in clinical narratives [3], [4], [5], [6]. Transformer encoders adapted to biomedical or clinical domains, including BioBERT, ClinicalBERT, PubMedBERT, and GatorTron, further improved task-specific extraction and relation modeling [7], [8], [9], [10], [11]. At the same time, large language models have made zero-shot or few-shot structured extraction from clinical notes feasible, but recent evaluations continue to report variability in strict format adherence, task-specific accuracy, and governance requirements [12], [13].
This work studies a narrower but operationally important question: for follow-up instruction extraction, is it better to ask a generative model to directly output action-date JSON, or to decompose the task into learned semantic extraction and deterministic date normalization? The repository evidence suggests that the decomposed design is promising on a controlled synthetic benchmark. The paper's central claim is therefore intentionally scoped: on synthetic, stress-varied outpatient notes, separating semantic span/link extraction from calendar arithmetic improves action-date reliability compared with the included direct generative baselines.
Contributions
- We define follow-up instruction extraction as a structured action-date extraction task over clinical notes, anchored by the visit date.
- We present a hybrid architecture with a shared BioBERT encoder, action/time BIO tagging, action-time span linking, and deterministic date normalization.
- We construct and evaluate on a 2,000-note synthetic outpatient corpus with five specialties, plan-section variation, 0/1/2 action cases, temporal shorthand, and distractors.
- We compare the hybrid pipeline against direct generative extraction baselines using span, linking, action-date, exact date, and date-error metrics.
- We identify the current evidence gaps that must be addressed before claims of real-world clinical robustness or deployment readiness are warranted.
Scope boundary
This draft does not claim clinical deployment readiness, real-note generalization, or superiority to all LLM prompting or fine-tuning strategies. The current evidence supports a controlled proof of concept. A top-tier submission will require external validation, ablations, confidence intervals, raw prediction release, and a reproducible experiment package.
2. Related Work
2.1 Clinical Information Extraction
Clinical information extraction aims to convert unstructured narrative text into structured representations usable for research and care operations. A broad review by Wang et al. describes clinical IE systems as pipelines that may include tokenization, syntactic processing, named entity recognition, concept identification, relation extraction, and context handling [1]. cTAKES is a prominent open-source clinical NLP system that extracts clinical concepts and attributes from free-text notes using a modular architecture [2]. Such systems demonstrate the long-standing value of task-specialized clinical NLP, but they usually target broad clinical concepts rather than executable follow-up action-date pairs.
The present task differs from standard concept extraction in two ways. First, the action to be scheduled is often a procedure, test, consult, or behavioral instruction rather than a diagnosis or medication mention. Second, extracting the action alone is insufficient: the system must link it to the temporal expression that determines when the action should occur. This places the task at the intersection of named entity recognition, relation extraction, and temporal normalization.
2.2 Temporal Information Extraction in Clinical Text
Clinical temporal extraction has been studied through shared tasks and corpora. The 2012 i2b2 challenge evaluated temporal relations in clinical text, including events, time expressions, and temporal links [3]. The THYME project developed clinical temporal annotation guidelines and corpora, emphasizing the specific demands of clinical narratives such as document sections, historical mentions, and event anchoring [4]. Clinical TempEval extended this line through SemEval tasks that evaluated systems on clinical temporal expression and relation extraction [5], [6].
These resources focus on general temporal structure in clinical narratives. Our task is narrower and more action-oriented: identify future follow-up actions and normalize their execution dates relative to a visit date. This narrower target is clinically useful but also creates new evaluation requirements. A model may correctly detect a temporal phrase and still fail if it links that phrase to the wrong action or if it produces the wrong calendar date.
2.3 Temporal Normalization and Rule-Based Date Handling
Temporal expression recognition and normalization have long used rule-based and hybrid systems. HeidelTime is a rule-based temporal tagger for extracting and normalizing temporal expressions [14], and SUTime provides a library for recognizing and normalizing time expressions using compositional rules [15]. Clinical temporal extraction work also emphasizes normalization as a foundational step for downstream temporal reasoning [16]. In this repository, deterministic date normalization is implemented with the Python dateparser library, using the encounter date as the relative base.
The key design choice is to avoid asking the neural model to do date arithmetic directly. Instead, the neural components identify spans and links, while deterministic logic maps phrases such as "within five weeks" or "11 mos" to ISO dates. This follows a broader reliability-first pattern: reserve learned models for ambiguous semantic interpretation and use deterministic modules for arithmetic or schema-constrained transformations whenever possible.
2.4 Domain-Specific Biomedical and Clinical Encoders
BERT established a general pretraining and fine-tuning framework for many NLP tasks [7]. Domain-specific pretraining then became important in biomedical and clinical NLP. BioBERT continues pretraining on PubMed abstracts and PMC full text and improves biomedical NER, relation extraction, and question answering [8]. ClinicalBERT adapts BERT to clinical notes and demonstrates the value of clinical-domain text for modeling EHR narratives [9]. PubMedBERT trains from scratch on biomedical literature and argues that in-domain vocabulary and pretraining matter for biomedical tasks [10]. GatorTron scales clinical language modeling using a large corpus of de-identified clinical text and evaluates across clinical NLP tasks including concept extraction and relation extraction [11].
Our use of BioBERT is therefore conservative rather than novel by itself. The novelty is not the encoder; it is the task-specific decomposition into action spans, time spans, action-time links, and deterministic date normalization for follow-up instructions.
2.5 Span-Based and Biaffine Relation Modeling
Relation extraction often benefits from explicit span-pair modeling. SpERT frames joint entity and relation extraction as a span-based transformer model, scoring candidate entities and relations over span representations [17]. Biaffine scoring, popularized in neural dependency parsing, provides a simple and effective way to score directed relations between two contextual representations [18]. The present architecture follows this family of ideas by representing action and time spans and scoring candidate action-time links, including a none option for actions without an explicit time.
2.6 LLM-Based Clinical Structured Extraction
Large language models have changed the baseline landscape for clinical information extraction. Agrawal et al. showed that large language models can act as zero- and few-shot clinical information extractors, introducing benchmark tasks based on clinical text [12]. Huang et al. assessed ChatGPT for structured extraction from clinical notes and reported promising feasibility, while also underscoring the need for careful evaluation against curated data [13]. Recent clinical foundation-model reviews emphasize schema adherence, privacy, governance, and evaluation designs that reflect health-system value rather than benchmark convenience [19]. Prompt-engineering studies further show that LLM extraction performance is sensitive to task instructions, annotation guidance, and few-shot examples [20].
These studies motivate using direct generative extraction as a serious baseline. However, follow-up action-date extraction places pressure on exact dates and linking correctness. The results in this repository suggest that direct generation can recover actions well but may suffer more on time extraction, linking, and date arithmetic. This makes the task a useful setting for comparing general-purpose generative extraction with task-specialized decomposed extraction.
2.7 Synthetic Clinical Data
Clinical NLP research is constrained by privacy, governance, and access barriers. Synthetic clinical text can reduce PHI risk and support controlled stress testing, but it does not replace real-world validation. Kweon et al. train a publicly shareable clinical LLM on synthetic clinical notes derived from public case reports and evaluate it against real-note tasks, illustrating one route for privacy-aware clinical NLP research [21]. A 2025 scoping review of synthetic health record generation highlights both promise and inconsistent evaluation practices across medical text, time series, and longitudinal data [22]. The present corpus should therefore be positioned as a controlled evaluation scaffold, not as evidence that the model is ready for real clinical deployment.
3. Task Definition
Each input example consists of a clinical note \(x\) and a visit date \(v\). The output is a set of follow-up items:
$$Y = \{(a_i, \tau_i, d_i)\}_{i=1}^{n},$$
where \(a_i\) is an action span, \(\tau_i\) is a time-expression span or null value, and \(d_i\) is an ISO-formatted execution date normalized relative to \(v\). The model must return an empty set when no scheduled follow-up action is present.
The task decomposes into four subtasks:
- Action span detection: identify text spans describing actionable future instructions, such as "MRI Brain", "X-Ray", or "Physical Therapy".
- Time span detection: identify temporal expressions that specify timing, such as "in 2 weeks", "within five weeks", or "11 mos".
- Action-time linking: assign each action to its corresponding time expression or to a none option.
- Date normalization: convert the linked time expression into an absolute date using the visit date as anchor.
The strict end-to-end criterion is set-level correctness over action-date pairs. A prediction is correct only when the action matches the gold action and the normalized date matches the gold date.
4. Dataset
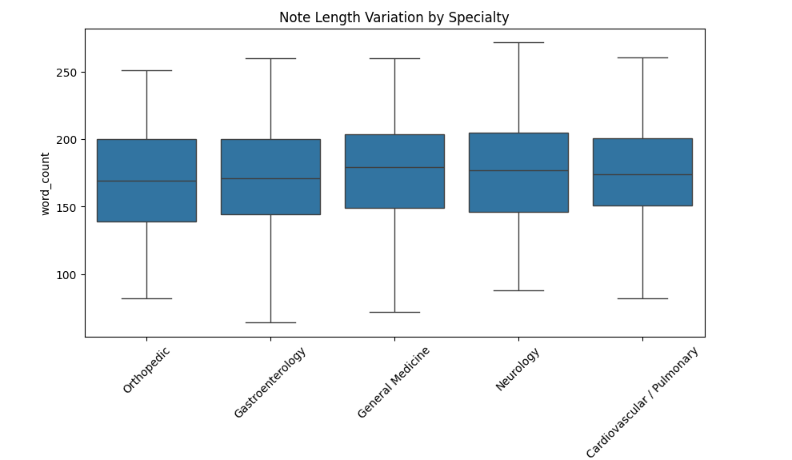
The repository contains Data/synthetic_clinical_notes_2000.csv, a synthetic corpus generated under a controlled schema to avoid use of protected health information. Each row contains the note text, visit date, specialty, topic, plan variant, action count, plan-section offsets, gold action/time spans, normalized dates, and style features.
| Property | Value |
|---|---|
| Total notes | 2,000 |
| Specialties | 5 |
| Orthopedic | 379 |
| Cardiovascular / Pulmonary | 391 |
| Gastroenterology | 394 |
| Neurology | 414 |
| General Medicine | 422 |
| 0-action notes | 497 |
| 1-action notes | 978 |
| 2-action notes | 525 |
| Mean actions per note | 1.014 |
| Note length range | 436-1,834 characters |
| Median note length | 1,193 characters |
| Recorded API errors | 0 |
| Recorded span errors | 26 |
The generator varies plan headers, specialty, clinical topic, clinician style features, action counts, and time-expression forms. Stress features include multi-action instructions, historical distractors, shorthand time phrases, proximity traps, and section ambiguity. The current draft treats these as controlled synthetic factors. A dataset card should be added before submission, including generation prompts, model version, sampling parameters, regeneration filters, annotation validation rules, and split identifiers.
5. Method
The proposed system uses a shared contextual encoder and two task heads. Let \(h_1,\ldots,h_T\) denote contextual token representations from BioBERT for a tokenized note window. Long notes are processed with sliding windows of maximum length 512 and document stride 128.
5.1 Span Tagging
The first head predicts BIO tags over tokens:
$$z_t \in \{\mathrm{O}, \mathrm{B\mbox{-}ACT}, \mathrm{I\mbox{-}ACT}, \mathrm{B\mbox{-}TIME}, \mathrm{I\mbox{-}TIME}\}.$$
The NER loss is weighted cross-entropy to reduce bias toward the majority O class:
$$\mathcal{L}_{\mathrm{NER}} = - \sum_{t=1}^{T} w_{z_t}\log p(z_t \mid h_t).$$
5.2 Span Representation
For each detected span \(s=(b,e)\), the model constructs a span representation using the start state, end state, and a width embedding:
$$r_s = [h_b; h_e; \phi(e-b+1)].$$
5.3 Action-Time Linking
For each action span \(a\) and candidate time span \(t\), the linker scores compatibility using a biaffine form with distance features:
$$\mathrm{score}(a,t) = r_a^\top W r_t + u^\top [r_a; r_t; \psi(\Delta_{a,t})] + b,$$
where \(\Delta_{a,t}\) encodes relative span distance. For each action, the model applies a softmax over all candidate time spans plus a none option. The linking loss is:
$$\mathcal{L}_{\mathrm{LINK}} = - \sum_{a \in A} \log p(t^\star_a \mid a, T \cup \{\varnothing\}).$$
5.4 Joint Objective
The total training objective is:
$$\mathcal{L} = \mathcal{L}_{\mathrm{NER}} + \alpha \mathcal{L}_{\mathrm{LINK}}, \quad \alpha = 1.$$
5.5 Date Normalization
After action-time linking, the linked time expression is normalized with deterministic date parsing. Given visit date \(v\) and time phrase \(\tau\), the date normalizer returns:
$$d = \mathrm{NormalizeDate}(\tau; v).$$
Implementation uses dateparser with RELATIVE_BASE set to the visit date and PREFER_DATES_FROM set to future. This design intentionally separates semantic extraction from calendar arithmetic.
6. Experiments
6.1 Models
The current repository compares three systems:
- BioBERT hybrid pipeline: the proposed NER, linking, and deterministic date-normalization system.
- ChatGPT zero-shot: direct JSON extraction from note text.
- LLaMA fine-tuned: an instruction model fine-tuned to output action-date JSON.
The baselines are useful, but a publication-quality experiment must preserve prompts, decoding parameters, model versions, checkpoint identifiers, train/validation/test split IDs, and raw predictions.
6.2 Metrics
We report exact span F1 for actions and times, linked-pair F1, action-date F1, exact date accuracy on matched actions, and mean absolute date error in days. Let \(P\) be the predicted set of action-date pairs and \(G\) the gold set. End-to-end precision, recall, and F1 are:
$$\mathrm{Precision} = \frac{|P \cap G|}{|P|}, \quad \mathrm{Recall} = \frac{|P \cap G|}{|G|}, \quad F_1 = \frac{2PR}{P+R}.$$
6.3 Current Results
| Model | Action F1 | Time/Date F1 | Action-Date F1 | Date Exact Acc. | Date MAE |
|---|---|---|---|---|---|
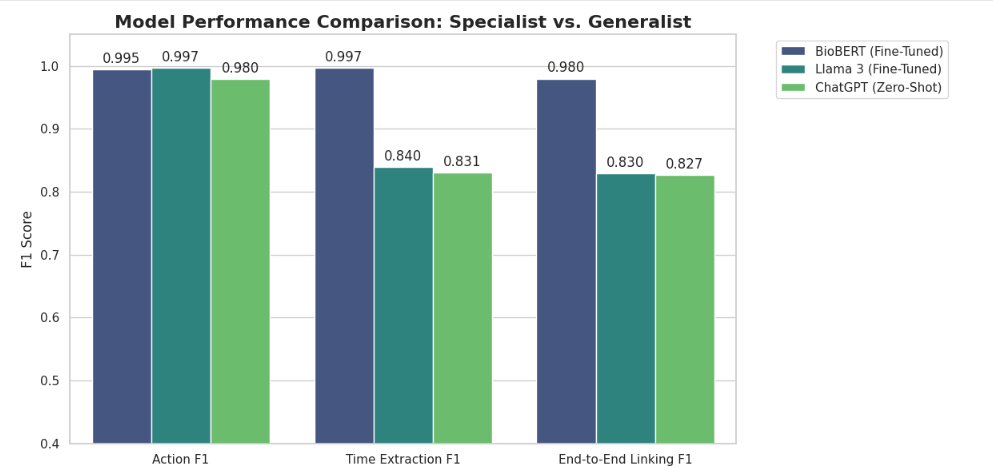
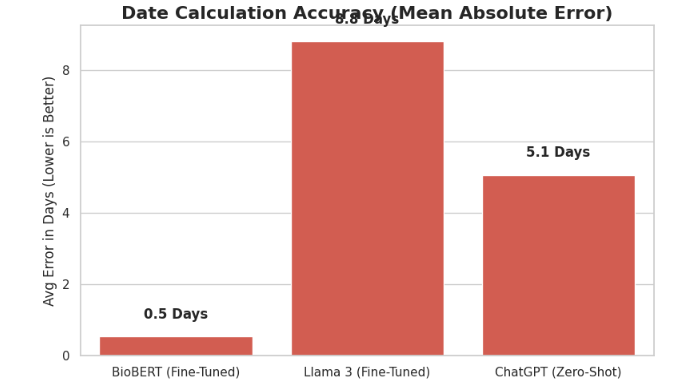
| BioBERT hybrid pipeline | 0.9949 | 0.9975 | 0.9797 | 0.9847 | 0.53 days |
| ChatGPT zero-shot | 0.9796 | 0.8308 | 0.8265 | 0.8438 | 5.07 days |
| LLaMA fine-tuned | 1.0000 | 0.8163 | 0.8061 | 0.8061 | 10.88 days |
6.4 Interpretation
The hybrid pipeline has high action and time span F1, and its action-date F1 is substantially higher than the included generative baselines. The largest observed gap is date arithmetic: the hybrid pipeline's mean absolute date error is below one day, while direct generative extraction produces larger errors. This supports the paper's central hypothesis that learned semantic extraction and deterministic date arithmetic should be separated for this task.
However, these results are not yet sufficient for a top-tier claim. They lack repeated seeds, confidence intervals, a documented split manifest, raw predictions, ablation studies, and external validation.
7. Required Ablations and Additional Analyses
A serious submission should add the following before claiming method superiority:
- Linking ablation: compare biaffine linking to nearest-time, same-sentence, action-order, and no-link baselines.
- Date-normalization ablation: compare deterministic normalization with direct model-generated dates while holding action/time extraction fixed.
- Encoder ablation: compare BioBERT with general BERT, PubMedBERT, ClinicalBERT, and a lightweight encoder if compute allows.
- Stress-slice analysis: report performance by specialty, note length, action count, shorthand expressions, historical distractors, proximity traps, and list-swapping cases.
- Uncertainty: use bootstrap confidence intervals over notes and, where feasible, repeated train/validation/test splits.
- Error taxonomy: manually classify errors into action boundary, time boundary, link selection, date normalization, no-action false positive, and missed instruction categories.
- External realism: evaluate on a small de-identified clinician-labeled note set or a public clinical temporal extraction benchmark adapted to follow-up action-date extraction.
8. Discussion
The results point to a useful design principle for clinical extraction systems: when the output combines semantic interpretation with exact symbolic transformation, reliability may improve when these functions are separated. The neural model is used to understand what textual spans denote actions and times and how they relate. The deterministic component is used to perform date arithmetic under a documented anchor date. This is aligned with clinical safety needs because date arithmetic errors are easier to audit and constrain in deterministic code than in unconstrained generation.
The comparison with LLM baselines should not be interpreted as a general indictment of generative models for clinical extraction. LLMs are flexible, can operate with limited task-specific labels, and may perform well with stronger prompts, tool use, constrained decoding, retrieval, or fine-tuning. The current evidence instead supports a narrower conclusion: for this synthetic follow-up extraction benchmark, a task-specific decomposed model outperforms the included direct JSON generation baselines on strict action-date correctness and date error.
The synthetic corpus is valuable for controlled stress testing because the true action, time, and date are known by construction. Yet synthetic data can encode generator artifacts, simplify real clinical variation, and overrepresent planned cases that fit the schema. Therefore, any top-tier version of this paper must include a realism bridge: either real-note evaluation, clinician annotation of a de-identified sample, or a rigorous synthetic-to-real validation study.
9. Limitations
- Synthetic-only evidence: the repository contains no real patient notes and cannot establish generalization to real EHR documentation.
- Single split: current metrics are point estimates on one held-out split, without confidence intervals or repeated seeds.
- Incomplete reproducibility: the main implementation is a Colab-style notebook combining generation, training, evaluation, and plotting.
- Baseline opacity: prompts, decoding settings, model versions, raw predictions, and split manifests are not yet fully preserved as auditable artifacts.
- Limited action ontology: the action set is specialty-specific and controlled; real instructions may include medications, referrals, conditional follow-up, patient behavior, and ambiguous responsibility.
- Dateparser assumptions: deterministic normalization depends on library behavior, locale assumptions, and phrase coverage.
- No workflow validation: the study does not evaluate clinician acceptance, EHR integration, safety monitoring, or downstream scheduling outcomes.
10. Ethics and Privacy
The current dataset is synthetic and includes no protected health information. This avoids direct privacy risk from patient records but does not remove the need for ethical review before real-world evaluation. Any extension to de-identified clinical notes should follow institutional governance, data-use agreements, annotation protocols, privacy review, and appropriate reporting of model failure modes. Because follow-up instructions can affect care coordination, deployment would require prospective validation, human oversight, alert-fatigue analysis, and clear accountability for scheduling decisions.
11. Reproducibility Plan
Before submission, the repository should be reorganized into a reproducible paper package:
scripts/generate_data.py: controlled data generation and validation.scripts/create_splits.py: deterministic split creation with saved note IDs.scripts/train_biobert.py: model training with config files and seeds.scripts/run_baselines.py: baseline inference with prompts and decoding parameters.scripts/evaluate.py: strict evaluation and bootstrap confidence intervals.scripts/make_figures.py: regenerate all manuscript figures and tables.predictions/: raw predictions for all systems.splits/: train/validation/test manifests.docs/dataset_card.mdanddocs/model_card.md: dataset and model documentation.
12. Conclusion
This draft presents a reliability-first approach to extracting follow-up actions and timelines from clinical notes. The core idea is to separate semantic understanding from date arithmetic: a BioBERT-based model detects action and time spans and links them, while deterministic normalization converts relative time phrases into absolute dates. On the current synthetic held-out benchmark, this decomposition yields stronger strict action-date performance and lower date error than the included direct generative baselines.
The work is promising but not yet complete as a top-tier paper. The next version should add ablations, stress-slice results, confidence intervals, raw prediction artifacts, and external validation. With those additions, the project can be framed as a focused clinical NLP methods paper about reliable action-date extraction rather than as a broad claim about clinical deployment or general LLM superiority.
Validated Bibliography
All entries below correspond to real papers or official project documentation verified through publisher pages, ACL Anthology, arXiv, PubMed/PMC, Nature, or project documentation during the literature search for this draft.
- Wang, Y., Wang, L., Rastegar-Mojarad, M., Moon, S., Shen, F., Afzal, N., Liu, S., Zeng, Y., Mehrabi, S., Sohn, S., and Liu, H. "Clinical information extraction applications: A literature review." Journal of Biomedical Informatics, 77, 34-49, 2018. DOI: 10.1016/j.jbi.2017.11.011.
- Savova, G. K., Masanz, J. J., Ogren, P. V., Zheng, J., Sohn, S., Kipper-Schuler, K. C., and Chute, C. G. "Mayo clinical Text Analysis and Knowledge Extraction System (cTAKES): architecture, component evaluation and applications." Journal of the American Medical Informatics Association, 17(5), 507-513, 2010. DOI: 10.1136/jamia.2009.001560.
- Sun, W., Rumshisky, A., and Uzuner, O. "Evaluating temporal relations in clinical text: 2012 i2b2 Challenge." Journal of the American Medical Informatics Association, 20(5), 806-813, 2013. DOI: 10.1136/amiajnl-2013-001628.
- Styler IV, W. F., Bethard, S., Finan, S., Palmer, M., Pradhan, S., de Groen, P. C., Erickson, B., Miller, T., Lin, C., Savova, G., and Pustejovsky, J. "Temporal Annotation in the Clinical Domain." Transactions of the Association for Computational Linguistics, 2, 143-154, 2014. DOI: 10.1162/tacl_a_00172.
- Bethard, S., Derczynski, L., Savova, G., Pustejovsky, J., and Verhagen, M. "SemEval-2015 Task 6: Clinical TempEval." Proceedings of the 9th International Workshop on Semantic Evaluation, 806-814, 2015. DOI: 10.18653/v1/S15-2136.
- Bethard, S., Savova, G., Palmer, M., and Pustejovsky, J. "SemEval-2016 Task 12: Clinical TempEval." Proceedings of the 10th International Workshop on Semantic Evaluation, 1052-1062, 2016. DOI: 10.18653/v1/S16-1165.
- Devlin, J., Chang, M.-W., Lee, K., and Toutanova, K. "BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding." Proceedings of NAACL-HLT, 4171-4186, 2019. DOI: 10.18653/v1/N19-1423.
- Lee, J., Yoon, W., Kim, S., Kim, D., Kim, S., So, C. H., and Kang, J. "BioBERT: a pre-trained biomedical language representation model for biomedical text mining." Bioinformatics, 36(4), 1234-1240, 2020. DOI: 10.1093/bioinformatics/btz682.
- Alsentzer, E., Murphy, J. R., Boag, W., Weng, W.-H., Jin, D., Naumann, T., and McDermott, M. "Publicly Available Clinical BERT Embeddings." Proceedings of the 2nd Clinical Natural Language Processing Workshop, 72-78, 2019. DOI: 10.18653/v1/W19-1909.
- Gu, Y., Tinn, R., Cheng, H., Lucas, M., Usuyama, N., Liu, X., Naumann, T., Gao, J., and Poon, H. "Domain-Specific Language Model Pretraining for Biomedical Natural Language Processing." ACM Transactions on Computing for Healthcare, 3(1), Article 2, 2021. DOI: 10.1145/3458754.
- Yang, X., Chen, A., PourNejatian, N., Shin, H. C., Smith, K. E., Parisien, C., Compas, C., Martin, C., Flores, M. G., Zhang, Y., Magoc, T., Harle, C. A., Lipori, G., Mitchell, D. A., Hogan, W. R., Shenkman, E. A., Bian, J., and Wu, Y. "A large language model for electronic health records." npj Digital Medicine, 5, 194, 2022. DOI: 10.1038/s41746-022-00742-2.
- Agrawal, M., Hegselmann, S., Lang, H., Kim, Y., and Sontag, D. "Large Language Models are Few-Shot Clinical Information Extractors." Proceedings of EMNLP, 1998-2022, 2022. DOI: 10.18653/v1/2022.emnlp-main.130.
- Huang, J., Yang, D. M., Rong, R., Nezafati, K., Treager, C., Chi, Z., Wang, S., Cheng, X., Guo, Y., Klesse, L. J., Xiao, G., Xie, Y., and others. "A critical assessment of using ChatGPT for extracting structured data from clinical notes." npj Digital Medicine, 7, 106, 2024. DOI: 10.1038/s41746-024-01079-8.
- Strotgen, J., and Gertz, M. "HeidelTime: High Quality Rule-based Extraction and Normalization of Temporal Expressions." Proceedings of the 5th International Workshop on Semantic Evaluation, 321-324, 2010. ACL Anthology ID: S10-1071.
- Chang, A. X., and Manning, C. D. "SUTime: A Library for Recognizing and Normalizing Time Expressions." Proceedings of the Eighth International Conference on Language Resources and Evaluation, 3735-3740, 2012.
- Lin, C., Miller, T., Dligach, D., Bethard, S., and Savova, G. "A BERT-based universal model for both within- and cross-sentence clinical temporal relation extraction." Proceedings of the 2nd Clinical Natural Language Processing Workshop, 65-71, 2019. DOI: 10.18653/v1/W19-1908.
- Eberts, M., and Ulges, A. "Span-based Joint Entity and Relation Extraction with Transformer Pre-training." Proceedings of the 24th European Conference on Artificial Intelligence, 2020. DOI: 10.3233/FAIA200321.
- Dozat, T., and Manning, C. D. "Deep Biaffine Attention for Neural Dependency Parsing." International Conference on Learning Representations, 2017.
- Wornow, M., Xu, Y., Thapa, R., Patel, B., Steinberg, E., Fleming, S., Pfeffer, M. A., Fries, J. A., and Shah, N. H. "The shaky foundations of large language models and foundation models for electronic health records." npj Digital Medicine, 6, 135, 2023. DOI: 10.1038/s41746-023-00879-8.
- Hu, Y., Chen, Q., Du, J., Peng, X., Keloth, V. K., Zuo, X., Zhou, Y., Li, Z., Jiang, X., Lu, Z., Roberts, K., and Xu, H. "Improving large language models for clinical named entity recognition via prompt engineering." Journal of the American Medical Informatics Association, 31(9), 1812-1820, 2024. DOI: 10.1093/jamia/ocad259.
- Kweon, S., Kim, J., Kim, J., Im, S., Cho, E., Bae, S., Oh, J., Lee, G., Moon, J. H., You, S. C., Baek, S., Han, C. H., Jung, Y. B., Jo, Y., and Choi, E. "Publicly Shareable Clinical Large Language Model Built on Synthetic Clinical Notes." Findings of the Association for Computational Linguistics: ACL 2024, 5148-5168, 2024. ACL Anthology ID: 2024.findings-acl.305.
- Loni, M., Poursalim, F., Asadi, M., and Gharehbaghi, A. "A review on generative AI models for synthetic medical text, time series, and longitudinal data." npj Digital Medicine, 8, 281, 2025. DOI: 10.1038/s41746-024-01409-w.